DISSCO
About the project
DiSSCo is a Research Infrastructure (RI) designed to manage, provide access to, and utilise both physical and digital natural science collections across Europe. It aims to unify these assets digitally across countries, ensuring data compliance with FAIR principles in the form of digital specimens representing FAIR digital objects. By bringing together 175 institutions from 23 countries, DiSSCo transforms fragmented resources into a cohesive knowledge base. It provides services for global research access and support, enhancing both physical and digital collection accessibility. By unlocking collections and integrating data, DiSSCo is accelerating research and supports novel research opportunities and collaborations.
The DiSSCo Flanders project “Towards a collection management infrastructure for Flanders”(2020-2024) involves 9 funded partners (MeiseBG, UGent, KU Leuven, VLIZ, ILVO, INBO, Royal Zoological Society of Antwerp, UAntwerp, VUB) and 5 associated partners (V.B.T.A., RBINS, RMCA, ULB, UNamur). The project aims to enhance the visibility and use of collections housed in Flemish institutions, ensuring their data and media are more Findable, Accessible, Interoperable, and Reusable (FAIR) (Trekels et al. 2022). It focuses on biological, anthropological, paleontological and geological collections, including preserved, living, tissue, and molecular samples, as well as smaller lab and orphan collections, to ensure their proper conservation and reusability. The project addresses the entire workflow, from specimen sampling in the field to physical collection management, databasing, digitization, and online publication.
IDLab role
Automated Preprocessing
Automated preprocessing methods based on computer vision have been developed to streamline the digitization of herbarium specimens. These methods begin by extracting the page boundary and color card from photographed specimens, which are then morphologically corrected and merged into a single normalised image. This approach significantly improved the digitization efficiency. We applied this method to historical herbarium collections from the late 18th and early 19th centuries (Charles Van Hoorebeke, Aimé Mac Leod, and Julius Mac Leod) as well as the collection of the Ghent University Museum (GUM).
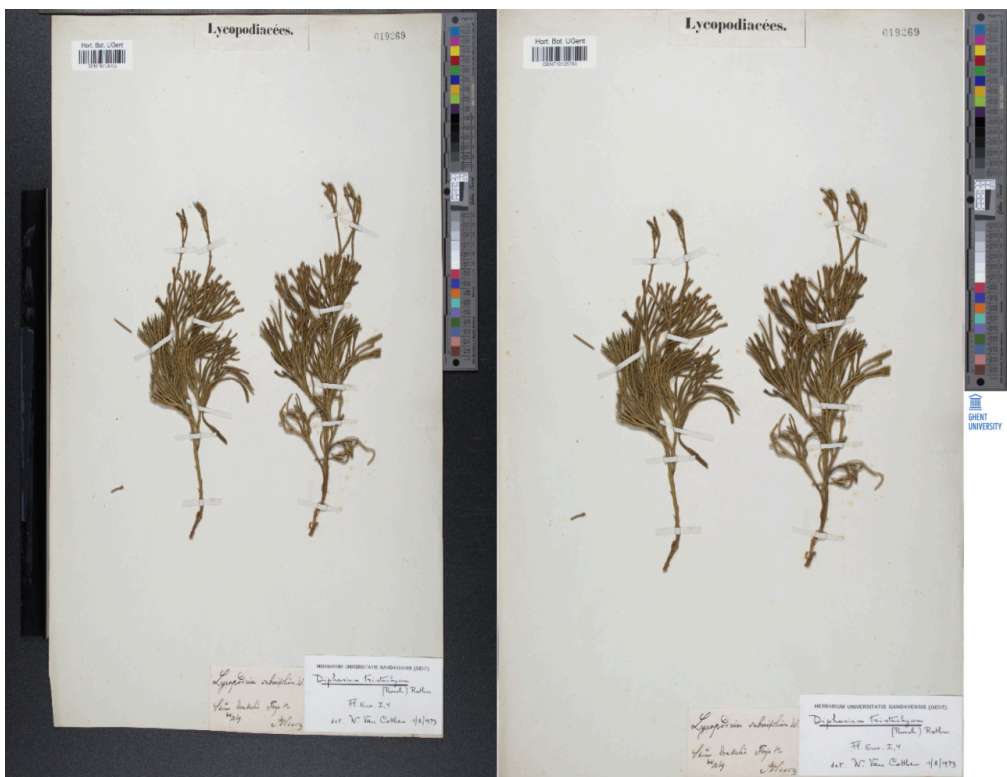

Metadata Extraction and linking
In the initial stage of DiSSCo, the primary focus was on extracting and identifying individual specimens from multi-specimen herbarium sheets (Thirukokaranam Chandrasekar 2021, Milleville et al. 2023). This process relied on recognizing the text present on the page for identification and association. Research into cross-collection linking was also performed, building upon the initial results from the FloreDeGand project (Thirukokaranam Chandrasekar et al. 2021). This allowed for connections to be established between herbarium specimens and related collections, such as paintings, field observations, and historical documents, providing a more comprehensive view of botanical data across collections.
Next, research was performed to segment entire herbarium sheets using deep learning models. A semi-automated labelling method was developed, to significantly speed up the annotation of herbarium specimen masks. This tool was used to annotate a dataset (in COCO format) of 250 herbarium sheets, each containing masks not only for the specimens themselves but also for common herbarium objects (rulers, colour bars, notes, stamps, and barcodes). Several state-of-the-art models were fine-tuned on this dataset, including binary, instance, and panoptic models. For binary specimen segmentation, our results were in line with related work, achieving an IoU score of 0.951. Furthermore, we found that popular instance segmentation models like YOLOv8 and Mask R-CNN struggled segmenting plant specimens accurately, but worked well for the non-specimen objects. We found that Mask2Former, the most recent model we tested, performed best overall. Further improvements were observed by combining a binary segmentation model for the specimens (UNet++) with YOLOv8 for the common herbarium objects. The code, fine-tuned models, and dataset are freely available at: https://github.com/kymillev/herbarium-segmentation.

Publications
-
Milleville, K. (2023). Unlocking the potential of digital archives via artificial intelligence. Ghent University. Faculty of Engineering and Architecture, Ghent, Belgium. http://hdl.handle.net/1854/LU-01HA1VDK26RX4CWJF6XBXA3B5F
-
Groom, Q., Dillen, M., Addink, W., Ariño, A. H. H., Bölling, C., Bonnet, P., … Gaikwad, J. (2023). Envisaging a global infrastructure to exploit the potential of digitised collections. BIODIVERSITY DATA JOURNAL, 11. https://doi.org/10.3897/bdj.11.e109439
-
Milleville, K., Thirukokaranam Chandrasekar, K. K., Van de Weghe, N., & Verstockt, S. (2023). Evaluating segmentation approaches on digitized herbarium specimens. In G. Bebis, G. Ghiasi, Y. Fang, A. Sharf, Y. Dong, C. Weaver, … L. Kohli (Eds.), ADVANCES IN VISUAL COMPUTING, ISVC 2023, PT II (Vol. 14362, pp. 65–78). https://doi.org/10.1007/978-3-031-47966-3_6
-
Milleville, K., Thirukokaranam Chandrasekar, K. K., & Verstockt, S. (2023). Automatic extraction of specimens from multi-specimen herbaria. ACM JOURNAL ON COMPUTING AND CULTURAL HERITAGE, 16(1). https://doi.org/10.1145/3575862
-
Thirukokaranam Chandrasekar, K. K. (2022). Meta data enrichment for improving the quality and usability of botanical collections. Ghent University. Faculty of Engineering and Architecture, Ghent, Belgium. http://hdl.handle.net/1854/LU-8760438
-
Thirukokaranam Chandrasekar, K. K., Milleville, K., & Verstockt, S. (2021). Species detection and segmentation of multi-specimen historical herbaria. BIODIVERSITY INFORMATION SCIENCE AND STANDARDS, 5. https://doi.org/10.3897/biss.5.74060
-
Thirukokaranam Chandrasekar, K. K., & Verstockt, S. (2020). Page boundary extraction of bound historical herbaria. In A. P. Rocha, L. Steels, & J. Van Den Herik (Eds.), ICAART: PROCEEDINGS OF THE 12TH INTERNATIONAL CONFERENCE ON AGENTS AND ARTIFICIAL INTELLIGENCE, VOL 1 (pp. 476–483). https://doi.org/10.5220/0009154104760483


